Getting started with R
Session 3 of 4: Data visualisation
Freddie Heather
Recap from session 2
Recap questions
- What is a script?
- Name five data manipulation functions from last week.
- What is the ‘pipe’ operator?
- What does “>=”, and “!=” do, and how would you use them?
- I want to create a new column in a tibble. What function would I use?
- I filter my data (e.g., age >= 3), name three way to check if the filter has worked?
- What is the ‘tidyverse’?
Today: Session 3
- Data import, checking, wrangling (recap)
- Data visualisation
- Critical thinking approach
Task
- Start a new R project, or open the project from last week.
- Create a new script and call it something informative (e.g.,
session3ordata_vis). - Comment your name, date, and a title at the top of the script.
- Save your script!
Data import & checking
Data import
Task
- Download the Reef Life Survey file ‘cape_howe.csv’ from: https://github.com/FreddieJH/r_workshop
- Import the csv and save it in R as an object (this will be your raw object and never overwritten)
- Create a new object that is only a single site - Choose one other than JBMP-S2
Data import
- Has it imported correctly?
- Sometimes you might have your hypothesis before data collection, not always
Task
- Use
head()and at least one other function to check to see things look okay. - How many columns? What do the columns mean?
- Think of a question(s) that you could answer using these data.
Data visualisation
Two numerical variables
- Hypothesis 1: I think that bigger fish are less abundant.
- What columns would I need to consider?
- How might I visualise this? boxplot? scatterplot? barplot?
A bit of wrangling first
- To answer the question if ‘bigger’ fish are more abundant, we are interested in the species, their mean size, and their total abundance.
Choosing the type of plot
| Predictor variable (x) | Response variable (y) | |
|---|---|---|
| Barplot, Boxplot, Violinplot | Categorical | Numeric |
| Scatterplot | Numeric | Numeric |
| Density, Histogram | Numeric | - |
Scatterplot using ggplot()
ggplot()is a function from theggplot2package (it’s part of thetidyversecollection of packages)- Scatterplot is used to compare two numerical variables
Mean size vs. total abundance
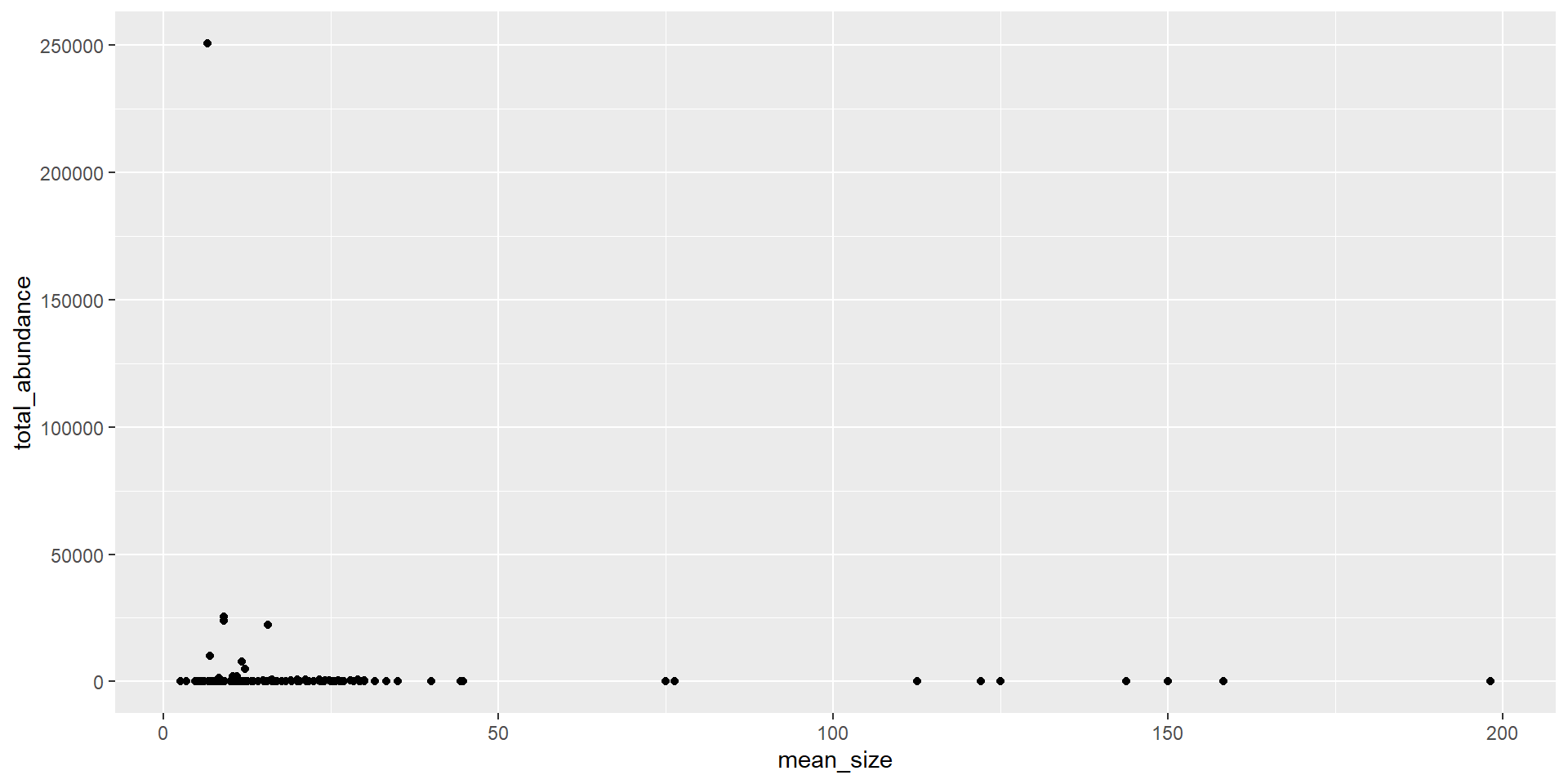
Transforming data
# Two ways transform data
# transform before input into the ggplot
sizes_byspp %>%
mutate(log_tot_abun = log(total_abundance)) %>%
ggplot(aes(x = mean_size, y = log_tot_abun)) +
geom_point()
# perform the transformation in the ggplot
sizes_byspp %>%
ggplot(aes(x = mean_size, y = total_abundance)) +
geom_point() +
scale_y_log10()
# what are the two differences between these two? (think: visual and data)Dealing with overalapping points
# Changing the transparency of the point (= alpha)
sizes_byspp %>%
ggplot(aes(x = mean_size, y = total_abundance)) +
geom_point(alpha = 0.5) + # alpha argument changes transparency
scale_y_log10()
# Changing the type of point (= pch)
sizes_byspp %>%
ggplot(aes(x = mean_size, y = total_abundance)) +
geom_point(pch = 21) + # pch argument changes point type (Google: "pch in r")
scale_y_log10()Task
- Change alpha to various values
- Change pch to get a hollow square
A cleaner plot
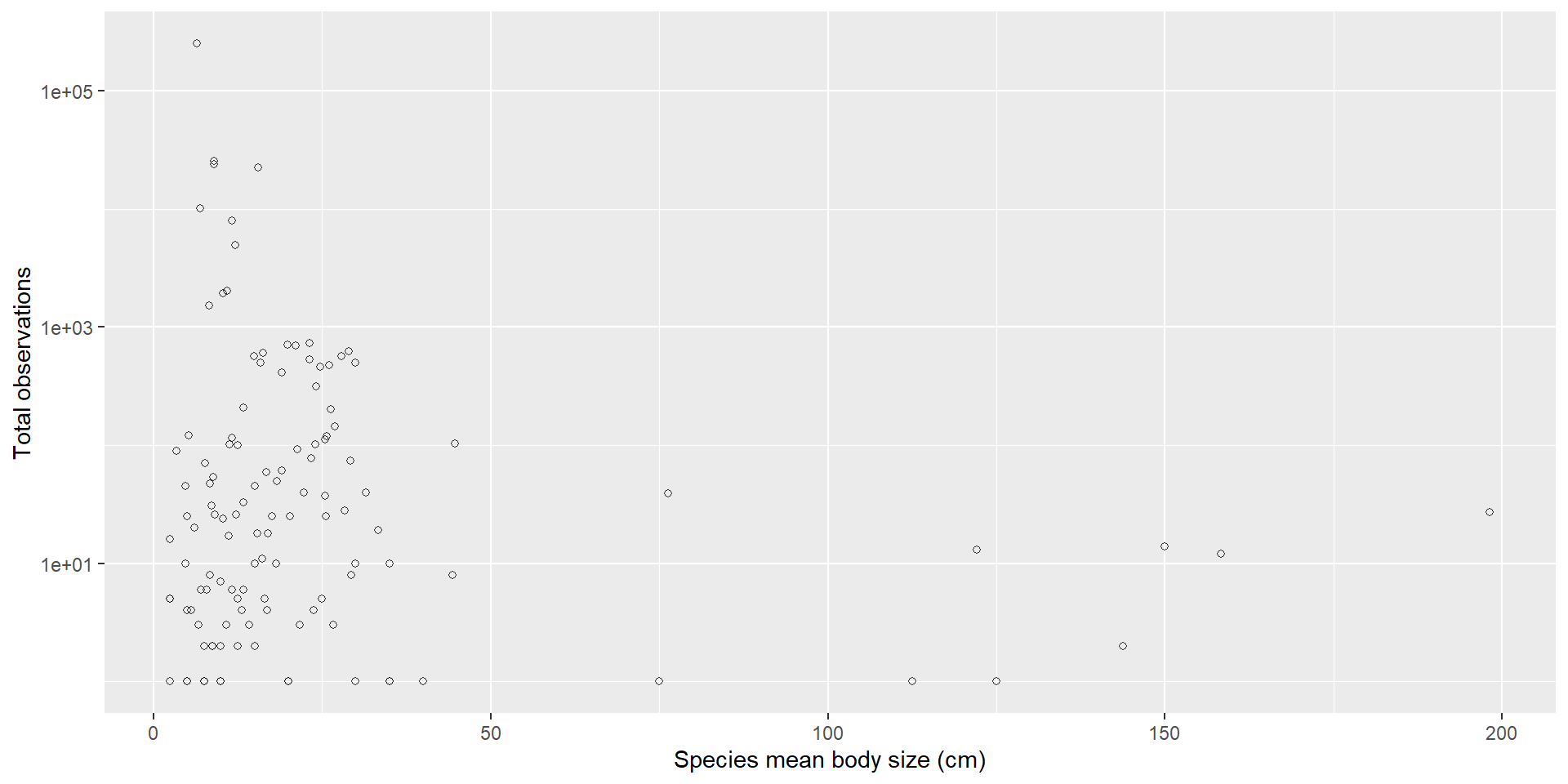
One numerical, one categorical
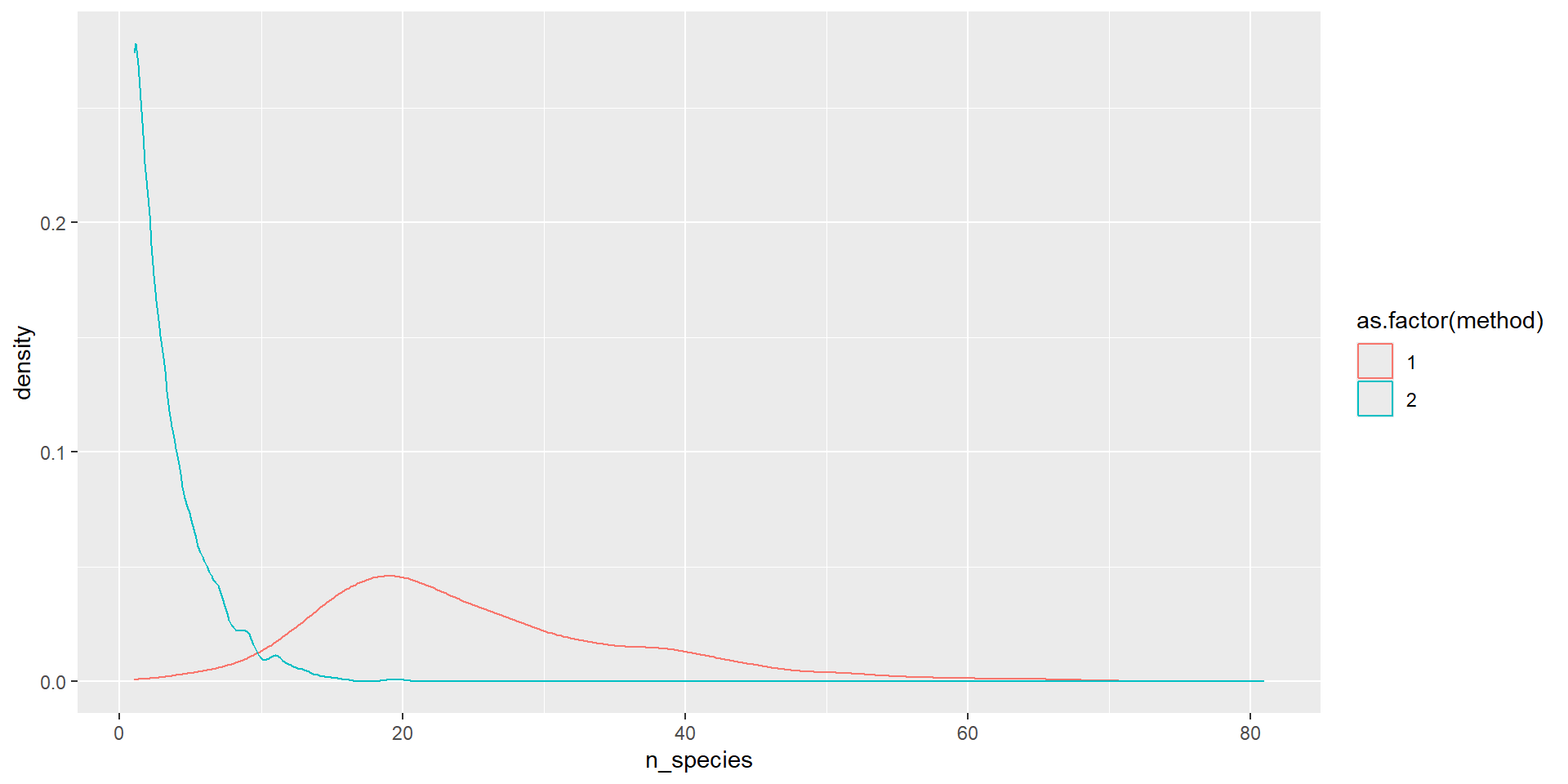
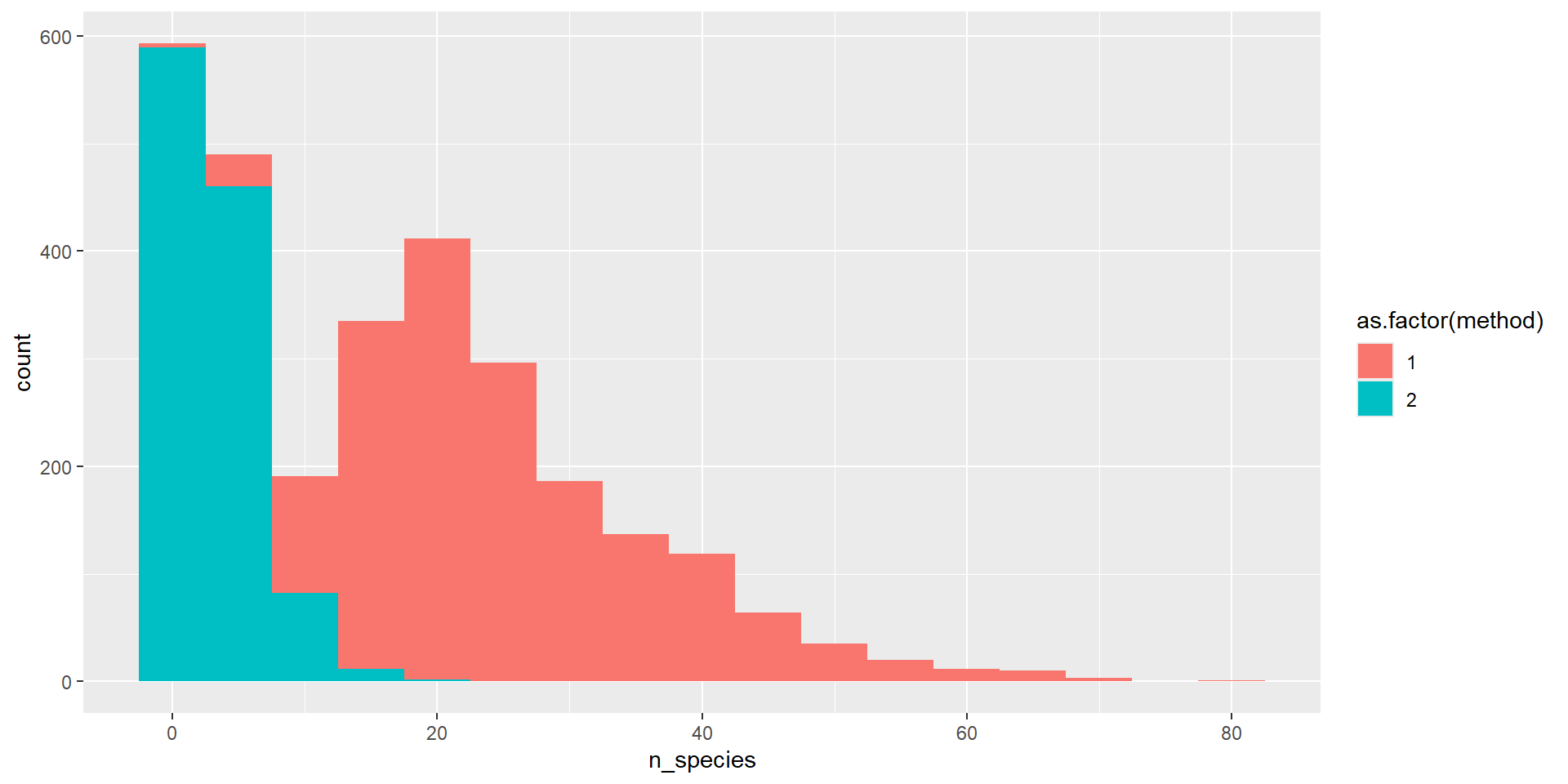
- Hypothesis 2: More species are seen in Method 1, than Method 2 (inverts + cryptic species)
- What columns do I need? What the classes of these columns?
- What would be suitable plots for these data?
Wrangling before plotting
Boxplot (Categorical vs numeric)
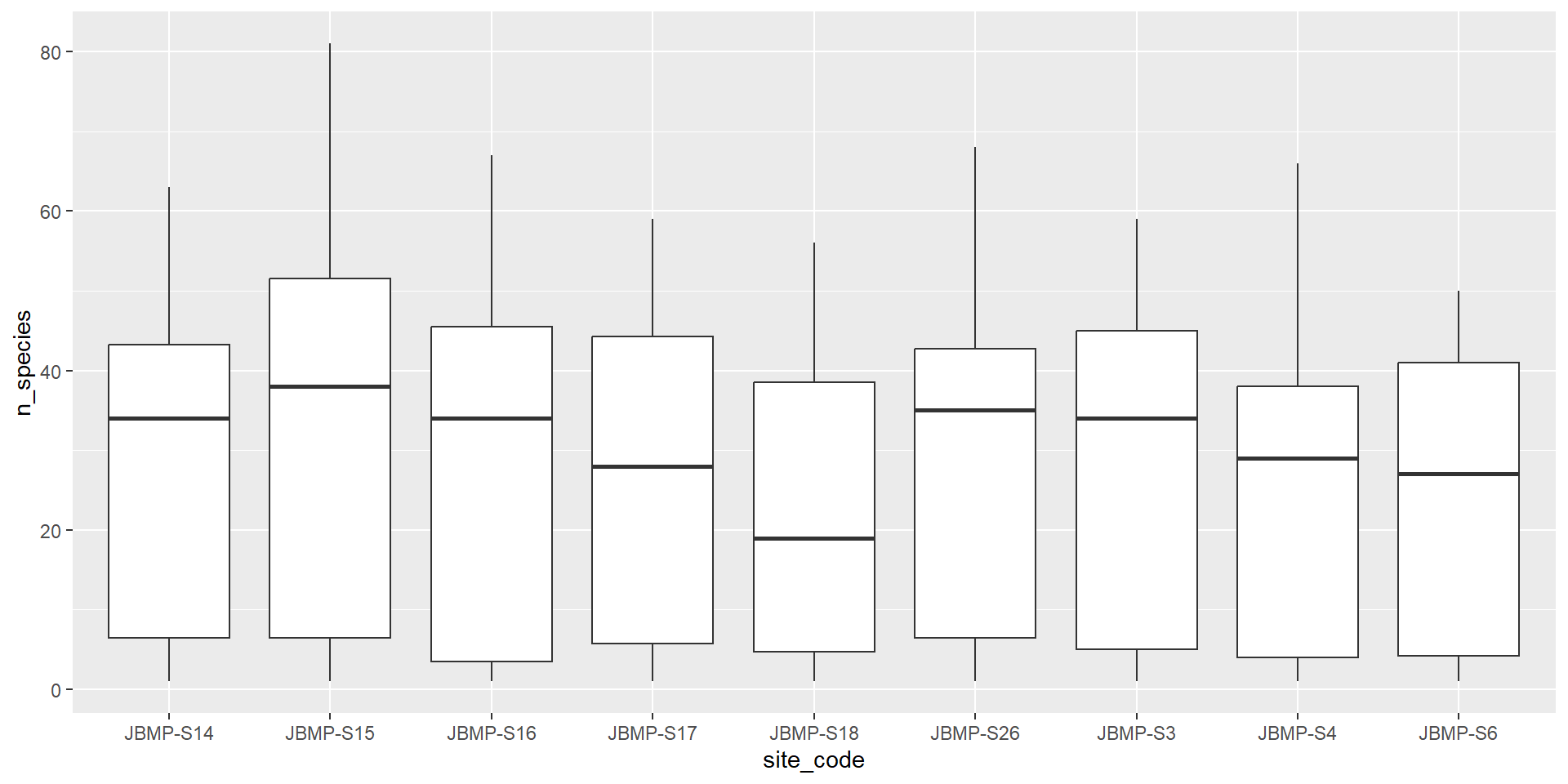
Boxplot coloured by method
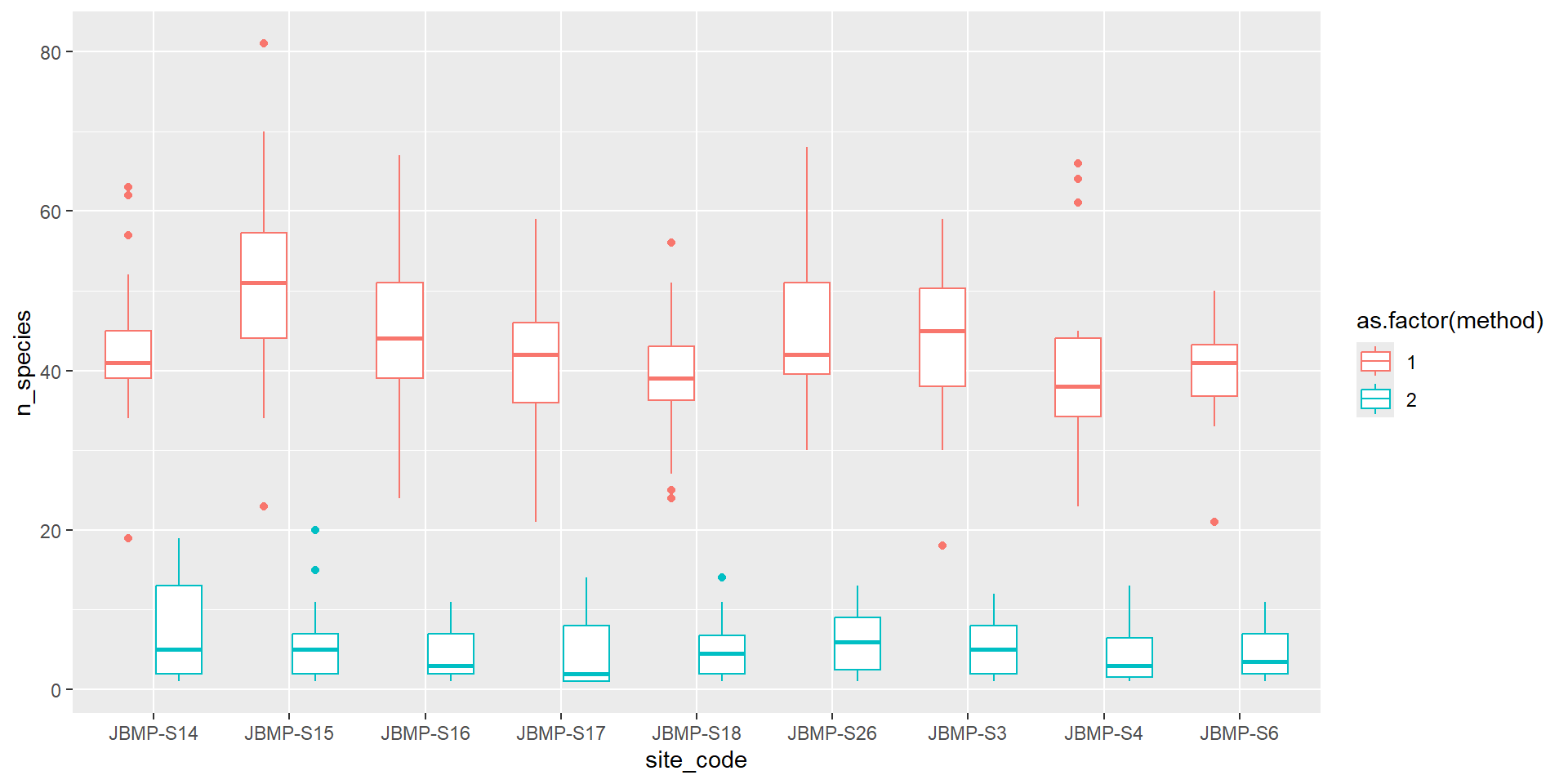
Violin plots (Categorical vs numeric)
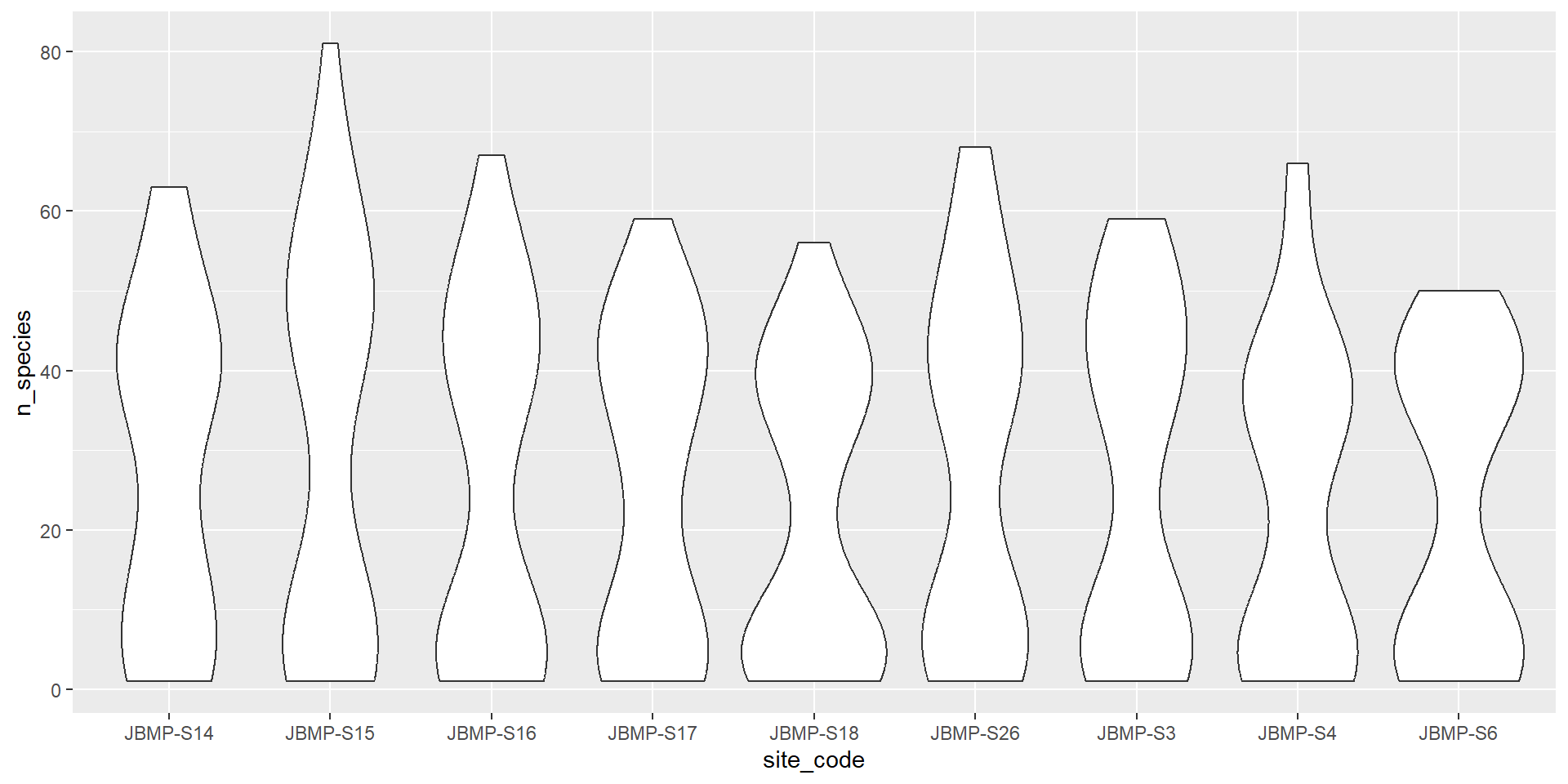
Barplots (Categorical vs numeric)
# For a barplot you might want the bar to represent the mean or median.
# how do barplots differ from violin or boxplots?
nspp_bysurv_bysite %>%
filter(site_code %in% selected_sites) %>%
summarise(
mean_diversity = mean(n_species, na.rm = TRUE),
.by = c(site_code, method)
) %>%
ggplot() +
aes(x = site_code, y = mean_diversity, fill = as.factor(method)) +
geom_col(position = "dodge") +
labs(fill = "Method", x = "Site Code", y = "Mean number of species")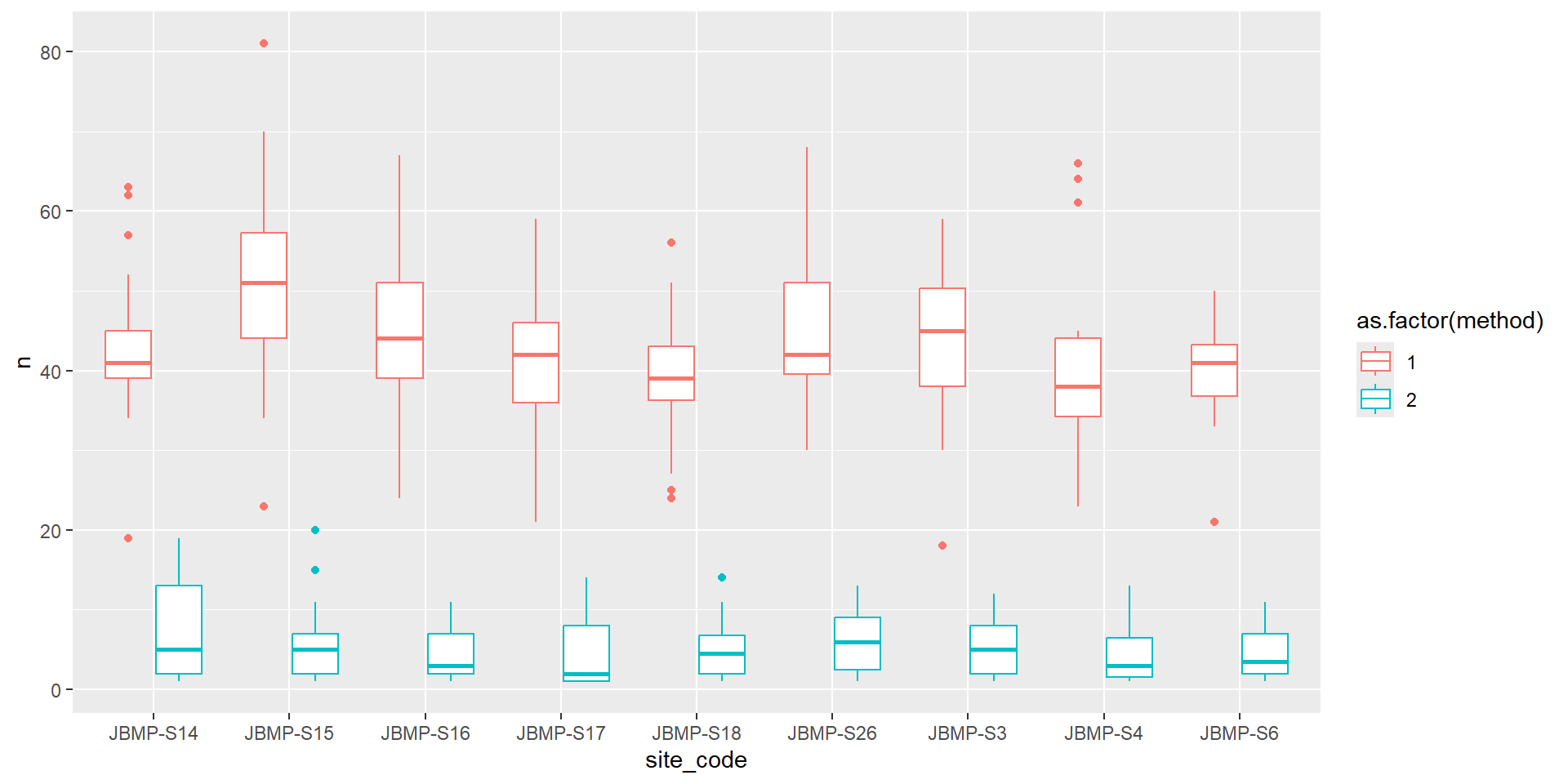
One numerical variable
Extra - multiple plots
- Sometime you want to split up plots by a factor
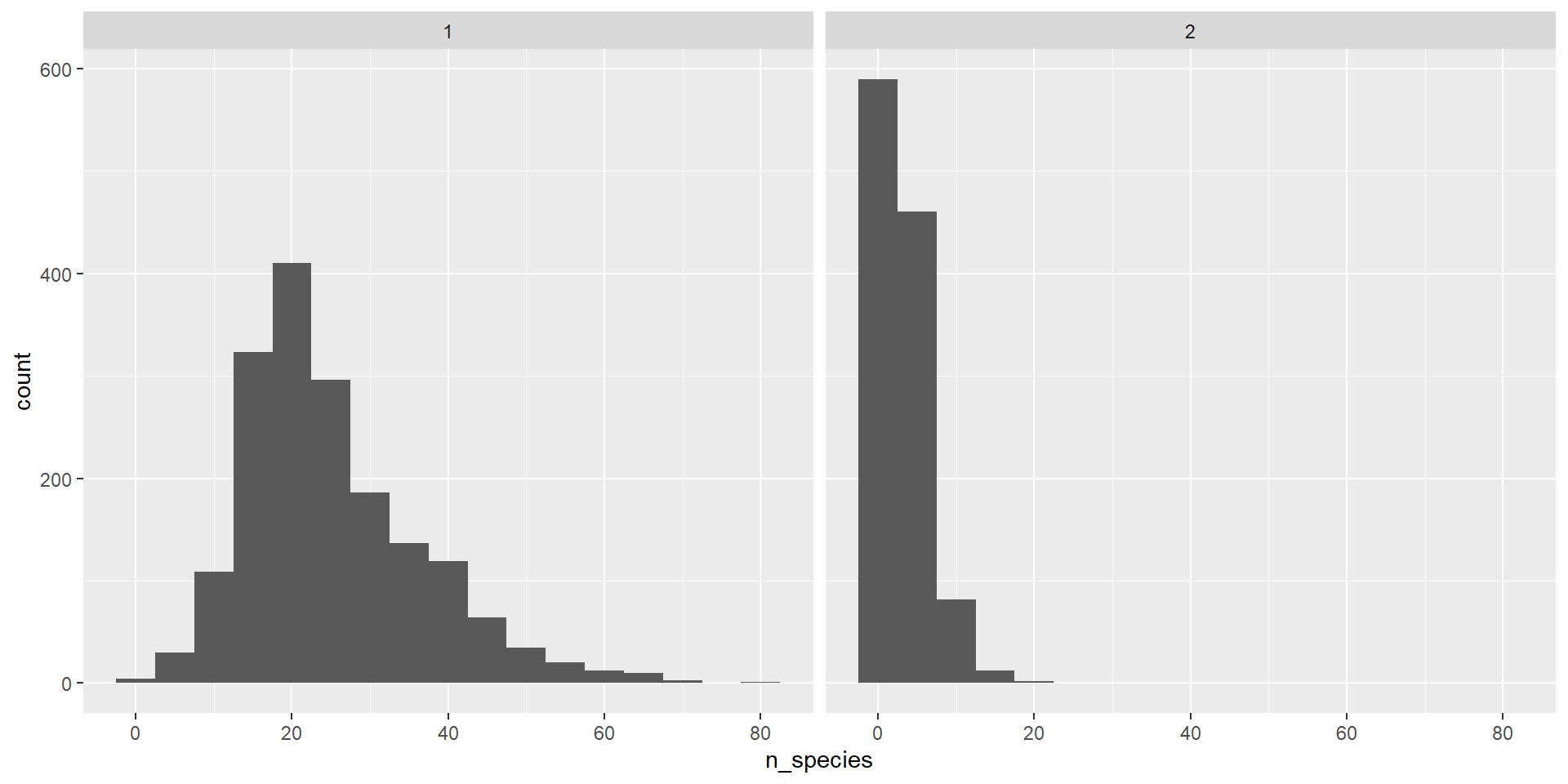
Extra - Error bars
# Barplots need error bars
nspp_bysurv_bysite %>%
filter(site_code %in% selected_sites) %>%
summarise(
mean_diversity = mean(n_species, na.rm = TRUE),
sd_diversity = sd(n_species, na.rm = TRUE),
.by = c(site_code, method)
) %>%
ggplot() +
aes(x = site_code, y = mean_diversity, fill = as.factor(method)) +
geom_col(position = position_dodge(width = 1)) +
geom_errorbar(
aes(
ymin = mean_diversity - sd_diversity,
ymax = mean_diversity + sd_diversity
),
width = 0.2,
position = position_dodge(width = 1)
) +
labs(fill = "Method", x = "Site Code", y = "Mean number of species")Extra - Theming
# Making pretty plots
nspp_bysurv_bysite %>%
filter(site_code %in% selected_sites) %>%
summarise(
mean_diversity = mean(n_species, na.rm = TRUE),
sd_diversity = sd(n_species, na.rm = TRUE),
.by = c(site_code, method)
) %>%
ggplot() +
aes(x = site_code, y = mean_diversity, fill = as.factor(method)) +
geom_col(position = position_dodge(width = 1)) +
geom_errorbar(
aes(
ymin = mean_diversity - sd_diversity,
ymax = mean_diversity + sd_diversity
),
width = 0.2,
position = position_dodge(width = 1)
) +
labs(fill = "Method", x = "Site Code", y = "Mean number of species") +
scale_fill_brewer(palette = "Set1") + #https://r-graph-gallery.com/38-rcolorbrewers-palettes.html
theme_classic() # https://ggplot2.tidyverse.org/reference/ggtheme.htmlExtra - Layering geoms
# Change the order of the geom_line() and geom_point()
nspp_bysurv_bysite %>%
mutate(year = year(survey_date)) %>%
summarise(mean_diversity = mean(n_species, na.rm = TRUE), .by = c(year, method)) %>%
ggplot() +
aes(x = year, y = mean_diversity) +
geom_line(aes(group = as.factor(method))) +
geom_point(aes(col = as.factor(method)), size = 5)Cheatsheet
Next week
- Collating all that we have learned.
- Please bring your own data if you have some.